from encodec.model import EncodecModel
import webdataset as wds
from whisperspeech.train import *
import pylab as plt
from IPython.display import Audio, HTML, displaySemantic to acoustic token modeling
Load the dataset
load_dataset
load_dataset (atoks_shard_spec:str, stoks_shard_dir:str, samples:int, random_trunc_p:float=0, vq_codes:int=4096, language:str='en', weight:float=1, validation:bool=False, exclude_files:str=None, randomize_speakers:bool=False, cwd:pathlib.Path=None)
| Type | Default | Details | |
|---|---|---|---|
| atoks_shard_spec | str | webdataset folder | |
| stoks_shard_dir | str | stoks webdataset base dir | |
| samples | int | samples per epoch | |
| random_trunc_p | float | 0 | probability of truncating the input to less than 30 seconds |
| vq_codes | int | 4096 | |
| language | str | en | |
| weight | float | 1 | |
| validation | bool | False | |
| exclude_files | str | None | |
| randomize_speakers | bool | False | |
| cwd | Path | None |
Model
import pylab as plt
import fastprogress
import IPython
import numpy as np
class CMLMVisual:
"""Visualize training progress"""
def __init__ (self, model, masterbar, total_steps):
self.model = model
self.masterbar = masterbar
self.total_steps = total_steps
self.epochs = total_steps // masterbar.main_bar.total
gs = plt.GridSpec(3, 1, height_ratios=[2,2,1])
graph_fig = plt.figure(figsize=(10,6))
self.graph_fig = graph_fig
self.loss_p = graph_fig.add_subplot(gs[0])
self.acc_p = graph_fig.add_subplot(gs[1], sharex=self.loss_p)
self.acc_p.tick_params('x', labelbottom=False)
self.lr_p = graph_fig.add_subplot(gs[2], sharex=self.loss_p)
self.lr_p.tick_params('x', labelbottom=False)
self.graph_out = None
self.its = []
self.train_losses = []
self.val_losses = []
self.lr_history = []
self.acc = np.nan
self.acc_history = []
self.pacc_history = []
def show(self):
self.start_t = time.time()
self.masterbar.write(["samples", "train", "val", "time"], table=True)
self.graph_out = display(self.graph_fig, display_id=True)
self.acc_out = display(IPython.display.HTML(''), display_id=True)
def hide(self):
if self.graph_out is not None:
self.graph_out.update(IPython.display.HTML(''))
def plot(self):
loss_p, acc_p, lr_p = self.loss_p, self.acc_p, self.lr_p
loss_p.clear()
loss_p.plot(self.its, self.train_losses)
loss_p.plot(self.its, self.val_losses)
loss_p.set_xlim(0, self.total_steps)
loss_p.set_yscale('log')
acc_p.clear()
for k in self.acc_history[-1].keys():
acc_p.plot(self.its, [x[k] for x in self.acc_history], ':')
lr_p.clear()
lrs = np.array(self.lr_history)
lr_p.plot(self.its, lrs)
self.graph_out.update(self.graph_fig)
def add_data(self, it, lr, train_loss, val_los):
self.its.append(it)
self.train_losses.append(train_loss)
self.val_losses.append(val_los)
self.lr_history.append(lr)
metrics = self.model.get_metrics()
self.acc_history.append(metrics)
html = "<h5>Accuracies:</h5><table>"
html += "<thead>"+(''.join([f"<td>{k}<td>" for k,x in metrics.items()]))+"</thead>"
html += "<tr>"+(''.join([f"<td>{x*100:.1f}%<td>" for k,x in metrics.items()]))+"</tr>"
html += "</table>"
self.acc_out.update(IPython.display.HTML(html))
self.plot()
def add_table_row(self, it, avg_train_loss, val_loss):
elapsed_t = time.time() - self.start_t
self.masterbar.write([it, f"{avg_train_loss:.5f}", f"{val_loss:.5f}", fastprogress.core.format_time(elapsed_t)], table=True)
def on_iter(self, bar, it, avg_train_loss, val_loss):
epoch = math.ceil(it / self.total_steps * self.epochs)
bar.comment = f"#{epoch}/{self.epochs} loss: {avg_train_loss:.3f} / {val_loss:.3f}"DelSumEmbedding
DelSumEmbedding (n_head=6, head_width=64, atoks_width=None, length=2250, codes=1024, quantizers=8, pos_embs=None)
Base class for all neural network modules.
Your models should also subclass this class.
Modules can also contain other Modules, allowing to nest them in a tree structure. You can assign the submodules as regular attributes::
import torch.nn as nn
import torch.nn.functional as F
class Model(nn.Module):
def __init__(self):
super().__init__()
self.conv1 = nn.Conv2d(1, 20, 5)
self.conv2 = nn.Conv2d(20, 20, 5)
def forward(self, x):
x = F.relu(self.conv1(x))
return F.relu(self.conv2(x))Submodules assigned in this way will be registered, and will have their parameters converted too when you call :meth:to, etc.
.. note:: As per the example above, an __init__() call to the parent class must be made before assignment on the child.
:ivar training: Boolean represents whether this module is in training or evaluation mode. :vartype training: bool
SADelARTransformer
SADelARTransformer (depth=3, ctx_n=2250, stoks_len=750, stoks_codes=4097, stoks_width=None, spk_width=None, atoks_width=None, n_head=3, head_width=64, ffn_mult=4, quantizers=8, speaker_map={'1': 0}, tunables=Tunables(init_std=9, embeddings_std=0.2, embeddings_lr_scale=10, output_mult=5.6, query_mult=0.3, encoder_depth_ratio=0.25, linear_heads=False, rope=True, q0_loss_mult=1, causal_encoder=False, lr0=0.003, clip_gradient_norm=2, weight_decay=0.001, warmup_steps=2000, random=False, random_finetune=False, force_hidden_to_emb=False))
Base class for all neural network modules.
Your models should also subclass this class.
Modules can also contain other Modules, allowing to nest them in a tree structure. You can assign the submodules as regular attributes::
import torch.nn as nn
import torch.nn.functional as F
class Model(nn.Module):
def __init__(self):
super().__init__()
self.conv1 = nn.Conv2d(1, 20, 5)
self.conv2 = nn.Conv2d(20, 20, 5)
def forward(self, x):
x = F.relu(self.conv1(x))
return F.relu(self.conv2(x))Submodules assigned in this way will be registered, and will have their parameters converted too when you call :meth:to, etc.
.. note:: As per the example above, an __init__() call to the parent class must be made before assignment on the child.
:ivar training: Boolean represents whether this module is in training or evaluation mode. :vartype training: bool
Tunables
Tunables (init_std:float=9, embeddings_std:float=0.2, embeddings_lr_scale:float=10, output_mult:float=5.6, query_mult:float=0.3, encoder_depth_ratio:float=0.25, linear_heads:bool=False, rope:bool=True, q0_loss_mult:float=1, causal_encoder:bool=False, lr0:float=0.003, clip_gradient_norm:float=2, weight_decay:float=0.001, warmup_steps:float=2000, random:bool=False, random_finetune:bool=False, force_hidden_to_emb:bool=False)
rand
rand (start, end)
DelSumHead
DelSumHead (quantizers=8, n_head=6, head_width=64)
Base class for all neural network modules.
Your models should also subclass this class.
Modules can also contain other Modules, allowing to nest them in a tree structure. You can assign the submodules as regular attributes::
import torch.nn as nn
import torch.nn.functional as F
class Model(nn.Module):
def __init__(self):
super().__init__()
self.conv1 = nn.Conv2d(1, 20, 5)
self.conv2 = nn.Conv2d(20, 20, 5)
def forward(self, x):
x = F.relu(self.conv1(x))
return F.relu(self.conv2(x))Submodules assigned in this way will be registered, and will have their parameters converted too when you call :meth:to, etc.
.. note:: As per the example above, an __init__() call to the parent class must be made before assignment on the child.
:ivar training: Boolean represents whether this module is in training or evaluation mode. :vartype training: bool
Training test
train_ds = load_dataset('../librilight/*atoks*.tar.gz', '../librilight-vq-en+pl/', 100000, vq_codes=513, exclude_files='../librilight/common-speakers-maxvad')
val_ds = load_dataset('../librilight/common-speakers-maxvad.tar.gz', '../librilight-vq-en+pl/', 512, vq_codes=513, validation=True)model = make_model('micro', quantizers=4, frozen_embeddings_model='vqmodel-medium-en+pl-512c-dim64.model',
tunables=Tunables()).cuda()
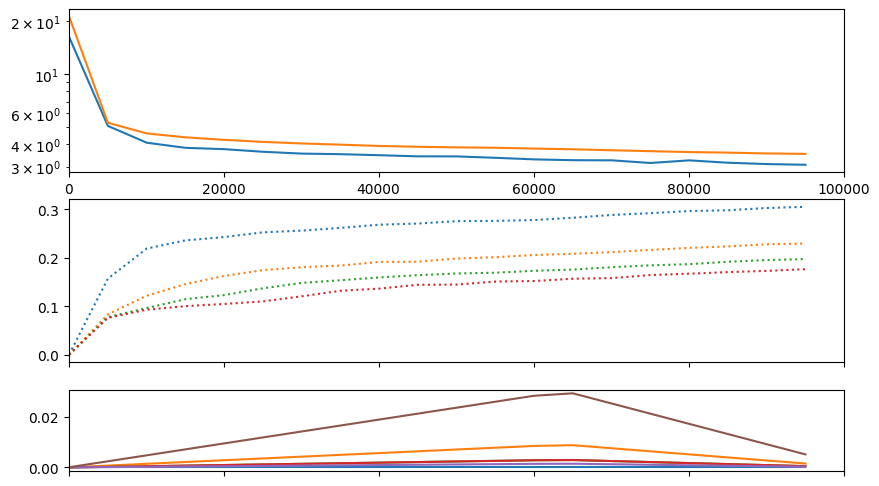
train(f"s2a-new", model, train_ds, val_ds, half=True, bs=32, lr=model.tunables.lr0, epochs=1, warmup_steps=model.tunables.warmup_steps,
table_row_every_iters=25000, run_valid_every_iters=5000, visual_class=CMLMVisual)Accuracies:
| acc_0 | acc_1 | acc_2 | acc_3 | ||||
| 29.6% | 23.6% | 21.2% | 19.2% |
| samples | train | val | time |
|---|---|---|---|
| 25024 | 3.95886 | 4.17079 | 02:34 |
| 50016 | 3.71909 | 3.81947 | 04:56 |
| 75008 | 3.53838 | 3.62924 | 07:18 |
| 100000 | 3.34118 | 3.46100 | 09:39 |
/opt/conda/lib/python3.10/site-packages/torch/optim/lr_scheduler.py:149: UserWarning: The epoch parameter in `scheduler.step()` was not necessary and is being deprecated where possible. Please use `scheduler.step()` to step the scheduler. During the deprecation, if epoch is different from None, the closed form is used instead of the new chainable form, where available. Please open an issue if you are unable to replicate your use case: https://github.com/pytorch/pytorch/issues/new/choose.
warnings.warn(EPOCH_DEPRECATION_WARNING, UserWarning)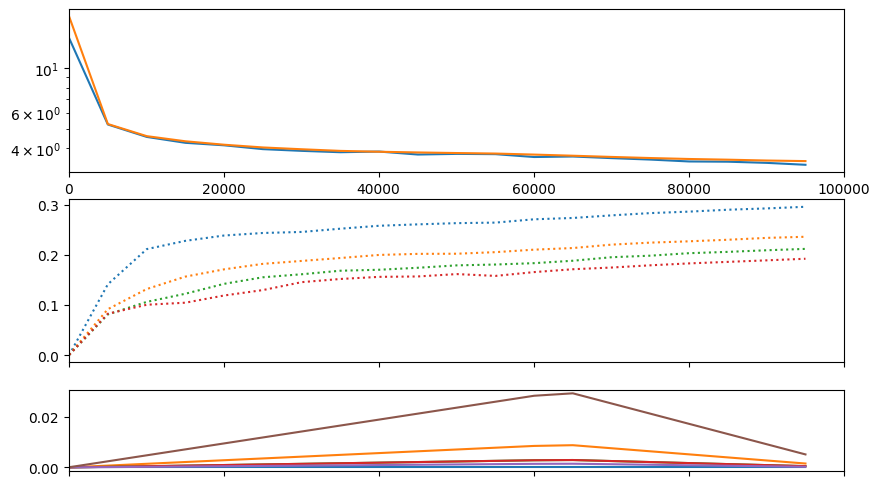
# encoder loss barely helps, probably because the RoPE cross-attention bias is already helping a lot
model = make_model('micro', quantizers=4, frozen_embeddings_model='vqmodel-medium-en+pl-512c-dim64.model',
tunables=Tunables(causal_encoder=True)).cuda()
train(f"s2a-new", model, train_ds, val_ds, half=True, bs=32, lr=model.tunables.lr0, epochs=1, warmup_steps=model.tunables.warmup_steps,
table_row_every_iters=25000, run_valid_every_iters=5000, visual_class=CMLMVisual)Accuracies:
| acc_0 | acc_1 | acc_2 | acc_3 | ||||
| 29.6% | 23.8% | 21.2% | 19.2% |
| samples | train | val | time |
|---|---|---|---|
| 25024 | 4.16333 | 4.16063 | 02:32 |
| 50016 | 3.98411 | 3.79632 | 04:55 |
| 75008 | 3.75278 | 3.62357 | 07:18 |
| 100000 | 3.54639 | 3.45734 | 09:41 |
/opt/conda/lib/python3.10/site-packages/torch/optim/lr_scheduler.py:149: UserWarning: The epoch parameter in `scheduler.step()` was not necessary and is being deprecated where possible. Please use `scheduler.step()` to step the scheduler. During the deprecation, if epoch is different from None, the closed form is used instead of the new chainable form, where available. Please open an issue if you are unable to replicate your use case: https://github.com/pytorch/pytorch/issues/new/choose.
warnings.warn(EPOCH_DEPRECATION_WARNING, UserWarning)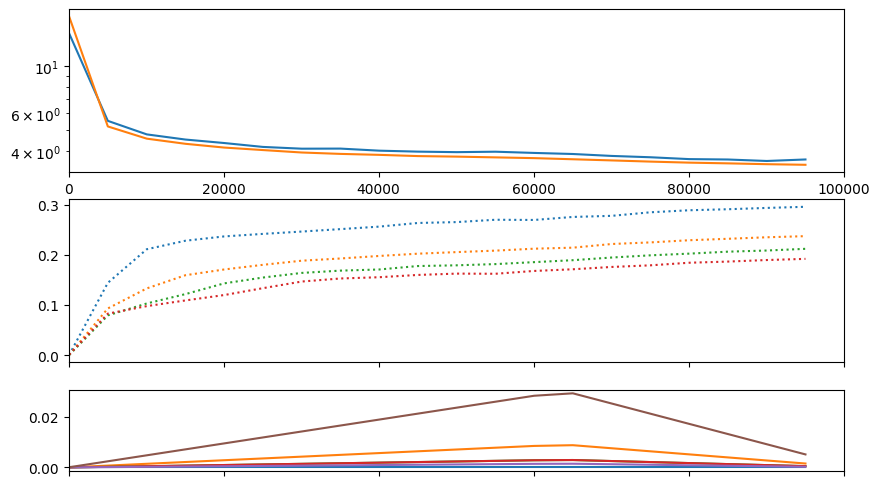
# we can prioritize the loss for the first quantizer
# we'd have to compare generations to really know if it helps though
model = make_model('micro', quantizers=4, frozen_embeddings_model='vqmodel-medium-en+pl-512c-dim64.model',
tunables=Tunables(q0_loss_mult=5)).cuda()
train(f"s2a-new", model, train_ds, val_ds, half=True, bs=32, lr=model.tunables.lr0, epochs=1, warmup_steps=model.tunables.warmup_steps,
table_row_every_iters=25000, run_valid_every_iters=5000, visual_class=CMLMVisual)Accuracies:
| acc_0 | acc_1 | acc_2 | acc_3 | ||||
| 30.5% | 23.0% | 19.8% | 17.7% |
| samples | train | val | time |
|---|---|---|---|
| 25024 | 3.59923 | 4.24838 | 02:32 |
| 50016 | 3.41711 | 3.88030 | 04:55 |
| 75008 | 3.19359 | 3.70881 | 07:17 |
| 100000 | 3.04986 | 3.53762 | 09:39 |
/opt/conda/lib/python3.10/site-packages/torch/optim/lr_scheduler.py:149: UserWarning: The epoch parameter in `scheduler.step()` was not necessary and is being deprecated where possible. Please use `scheduler.step()` to step the scheduler. During the deprecation, if epoch is different from None, the closed form is used instead of the new chainable form, where available. Please open an issue if you are unable to replicate your use case: https://github.com/pytorch/pytorch/issues/new/choose.
warnings.warn(EPOCH_DEPRECATION_WARNING, UserWarning)